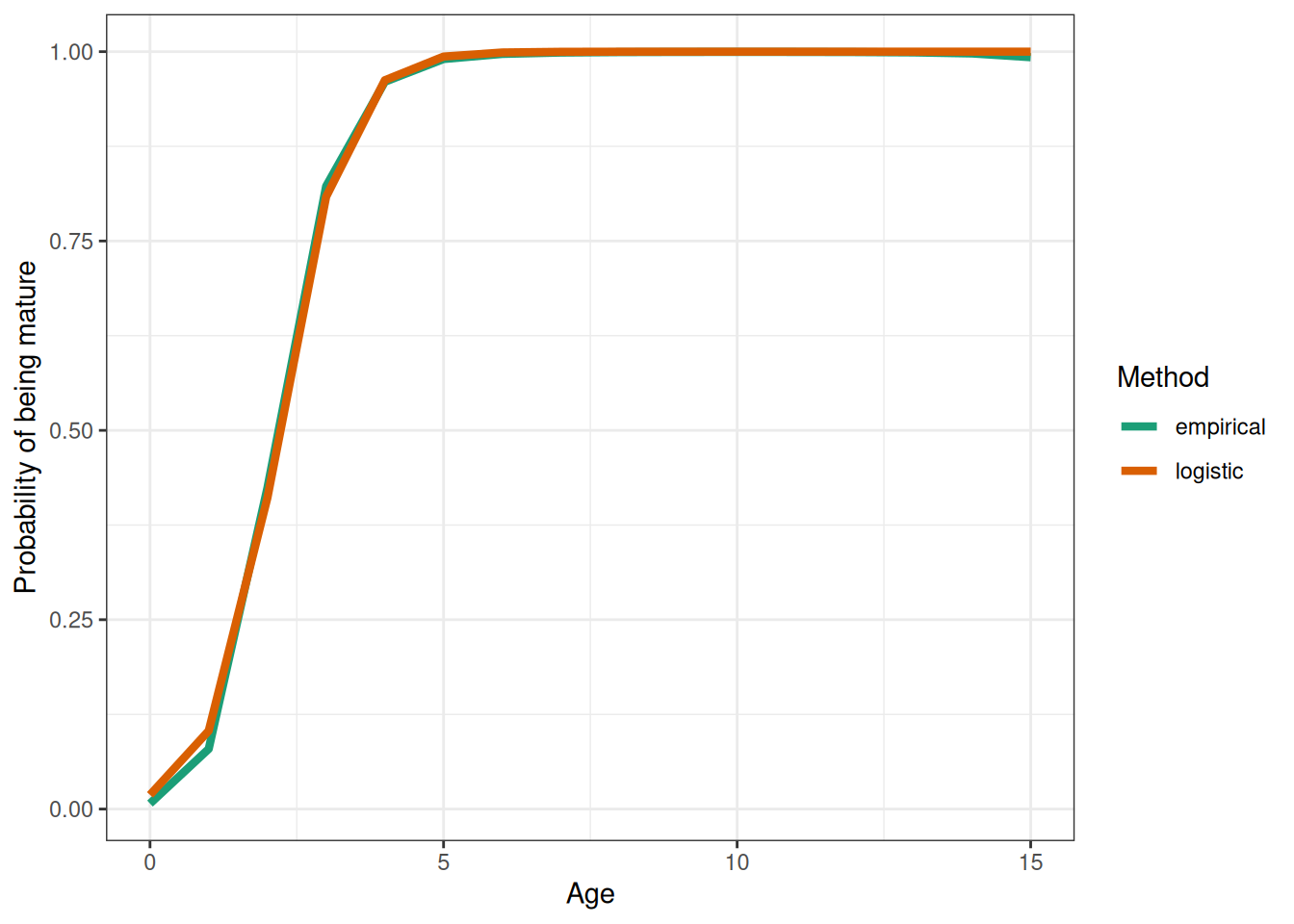
time_varying_selectivity <- TRUE
selectivity_used <- "Logistic"
FIMS::clear()
parameters <- FIMS::create_default_configurations(data = data_4_model) |>
tidyr::unnest(data) |>
dplyr::rows_update(
tibble::tibble(
module_name = "Selectivity",
fleet_name = c("Fishery", "Acoustic_Survey"),
module_type = c(selectivity_used, "Logistic")
),
by = c("module_name", "fleet_name")
) |>
FIMS::create_default_parameters(data = data_4_model) |>
tidyr::unnest(data) |>
dplyr::rows_update(
tibble::tibble(
module_name = c("Maturity", "Maturity"),
label = c("inflection_point", "slope"),
value = c(
hake_maturity_inflection_point,
hake_maturity_slope
)
),
by = c("module_name", "label")
) |>
dplyr::rows_update(
tibble::tibble(
module_name = c("Population"),
label = c("log_M"),
# estimation_type = "fixed_effects",
value = log(dplyr::pull(natural_mortality_matrix, m)),
time = dplyr::pull(natural_mortality_matrix, year),
age = dplyr::pull(natural_mortality_matrix, age)
),
by = c("module_name", "label", "time", "age")
) |>
dplyr::rows_update(
tibble::tibble(
module_name = "Population",
label = "log_init_naa",
age = FIMS::get_ages(data_4_model),
value = log(init_naa),
estimation_type = "fixed_effects"
),
by = c("module_name", "label", "age")
) |>
dplyr::rows_update(
tibble::tibble(
module_name = "Selectivity",
fleet_name = rep(c("Fishery", "Acoustic_Survey"), each = 2),
label = rep(c("inflection_point", "slope"), 2),
# TODO: play with these start values
value = c(2.05, 3.17, 4.4, 3.3),
estimation_type = "fixed_effects"
),
by = c("module_name", "fleet_name", "label")
) |>
dplyr::rows_update(
tibble::tibble(
module_name = "Fleet",
fleet_name = "Acoustic_Survey",
label = c("log_q"),
value = log(0.832),
estimation_type = "fixed_effects"
),
by = c("module_name", "fleet_name", "label")
)
fleet_names <- get_fleets(data_4_model)
# Initialize lists to store fleet-related objects
fleet <- fleet_selectivity <-
fleet_landings <- fleet_landings_distribution <-
fleet_index <- fleet_index_distribution <-
fleet_age_comp <- fleet_agecomp_distribution <-
vector("list", length(fleet_names))
for (i in seq_along(fleet_names)) {
# Selectivity
fleet_selectivity[[i]] <- FIMS:::initialize_selectivity(
parameters = parameters,
data = data_4_model,
fleet_name = fleet_names[i]
)
# Time-varying selectivity for the fishery
if (i == 1 & time_varying_selectivity) {
selectivity_type <- parameters |>
dplyr::filter(module_name == "Selectivity", fleet_name == "Fishery") |>
dplyr::slice(1) |>
dplyr::pull(module_type)
if (selectivity_type == "Logistic"){
fleet_selectivity[[i]]$inflection_point$resize(get_n_years(data_4_model))
for (ii in seq(get_n_years(data_4_model))) {
fleet_selectivity[[i]]$inflection_point[ii]$value <- 1
fleet_selectivity[[i]]$inflection_point[ii]$estimation_type$set("fixed_effects")
}
fleet_selectivity[[i]]$slope$resize(get_n_years(data_4_model))
for (ii in seq(get_n_years(data_4_model))) {
fleet_selectivity[[i]]$slope[ii]$value <- 1
fleet_selectivity[[i]]$slope[ii]$estimation_type$set("fixed_effects")
}
}
if (selectivity_type == "DoubleLogistic") {
fleet_selectivity[[i]]$inflection_point_asc$resize(get_n_years(data_4_model))
for (ii in seq(get_n_years(data_4_model))) {
fleet_selectivity[[i]]$inflection_point_asc[ii]$value <- 1
fleet_selectivity[[i]]$inflection_point_asc[ii]$estimation_type$set("fixed_effects")
}
# fleet_selectivity[[i]]$inflection_point_desc$resize(get_n_years(data_4_model))
# for (ii in seq(get_n_years(data_4_model))) {
# fleet_selectivity[[i]]$inflection_point_desc[ii]$value <- 1
# fleet_selectivity[[i]]$inflection_point_desc[ii]$estimation_type$set("fixed_effects")
# }
}
}
# Setup the fleets
fleet_module_ids <- c(
selectivity = fleet_selectivity[[i]]$get_id()
)
fleet_types <- get_data(data_4_model) |>
dplyr::filter(name == fleet_names[i]) |>
dplyr::pull(type) |>
unique()
data_distribution_names_for_fleet_i <- parameters |>
dplyr::filter(fleet_name == fleet_names[i] & distribution_type == "Data") |>
dplyr::pull(module_type)
# Landings
if ("landings" %in% fleet_types &&
"Landings" %in% data_distribution_names_for_fleet_i) {
# Initialize landings module for the current fleet
fleet_landings[[i]] <- FIMS:::initialize_landings(
data = data_4_model,
fleet_name = fleet_names[i]
)
# Add the module ID for the initialized landings to the list of fleet module IDs
fleet_module_ids <- c(
fleet_module_ids,
c(landings = fleet_landings[[i]]$get_id())
)
}
# Index
if ("index" %in% fleet_types &&
"Index" %in% data_distribution_names_for_fleet_i) {
# Initialize index module for the current fleet
fleet_index[[i]] <- FIMS:::initialize_index(
data = data_4_model,
fleet_name = fleet_names[i]
)
fleet_module_ids <- c(
fleet_module_ids,
c(index = fleet_index[[i]]$get_id())
)
}
# Age composition
if ("age_comp" %in% fleet_types &&
"AgeComp" %in% data_distribution_names_for_fleet_i) {
# Initialize age composition module for the current fleet
fleet_age_comp[[i]] <- FIMS:::initialize_comp(
data = data_4_model,
fleet_name = fleet_names[i],
type = "AgeComp"
)
fleet_module_ids <- c(
fleet_module_ids,
c(age_comp = fleet_age_comp[[i]]$get_id())
)
}
fleet[[i]] <- FIMS:::initialize_fleet(
parameters = parameters,
data = data_4_model,
fleet_name = fleet_names[i],
linked_ids = fleet_module_ids
)
# Fleet uncertainty
fleet_sd_input <- parameters |>
dplyr::filter(fleet_name == fleet_names[i] & label == "log_sd") |>
dplyr::mutate(
label = "sd",
value = exp(value)
)
if ("index" %in% fleet_types &&
"Index" %in% data_distribution_names_for_fleet_i) {
fleet_index_distribution[[i]] <- FIMS:::initialize_data_distribution(
module = fleet[[i]],
family = lognormal(link = "log"),
sd = fleet_sd_input,
data_type = "index"
)
}
if ("landings" %in% fleet_types &&
"Landings" %in% data_distribution_names_for_fleet_i) {
fleet_landings_distribution[[i]] <- FIMS:::initialize_data_distribution(
module = fleet[[i]],
family = lognormal(link = "log"),
sd = fleet_sd_input,
data_type = "landings"
)
}
if ("age_comp" %in% fleet_types &&
"AgeComp" %in% data_distribution_names_for_fleet_i) {
fleet_agecomp_distribution[[i]] <- FIMS:::initialize_data_distribution(
module = fleet[[i]],
family = multinomial(link = "logit"),
data_type = "agecomp"
)
}
}
# Recruitment
recruitment <- methods::new(BevertonHoltRecruitment)
recruitment_process <- methods::new(LogDevsRecruitmentProcess)
recruitment$SetRecruitmentProcessID(recruitment_process$get_id())
# set up log_rzero (equilibrium recruitment)
recruitment$log_rzero[1]$value <- inputs[["ctl"]][["SR_parms"]]["SR_LN(R0)", "INIT"]
recruitment$log_rzero[1]$estimation_type$set("fixed_effects")
# set up logit_steep
recruitment$logit_steep[1]$value <- FIMS::logit(0.2, 1.0, steepness)
recruitment$logit_steep[1]$estimation_type$set("constant")
# recruit deviations should enter the model in normal space.
# The log is taken in the likelihood calculations
# Added an element to the log_devs vector because we need recruitment in the
# terminal year + 1
recruitment$log_devs$resize(get_n_years(data_4_model))
for (y in seq(recruitment$log_devs$size())) {
recruitment$log_devs[y]$value <- 0
}
recruitment$log_devs$set_all_estimable(TRUE)
recruitment$n_years$set(get_n_years(data_4_model))
recruitment_distribution <- methods::new(DnormDistribution)
# TODO: check with Andrea about log space here
# logR_sd needs to enter the model logged b/c the exp() is
# taken before the likelihood calculation
recruitment_distribution$log_sd[1]$value <-
inputs[["ctl"]][["SR_parms"]]["SR_sigmaR", "INIT"]
# TODO: check with Andrea about this not being in the parameter vector
recruitment_distribution$log_sd[1]$estimation_type$set("fixed_effects")
recruitment_distribution$observed_values$resize(recruitment$log_devs$size())
recruitment_distribution$expected_values$resize(recruitment$log_devs$size())
for (i in seq(recruitment$log_devs$size())) {
recruitment_distribution$observed_values[i]$value <- 0
recruitment_distribution$expected_values[i]$value <- 0
}
recruitment_distribution$set_distribution_links(
"random_effects",
recruitment$log_devs$get_id()
)
recruitment$log_devs$set_all_random(TRUE)
# recruitment$log_devs[46]$estimation_type$set("constant")
# recruitment$log_devs[47]$estimation_type$set("constant")
# recruitment$log_devs[48]$estimation_type$set("constant")
# Put a prior on log_rzero
log_rzero_prior <- methods::new(DnormDistribution)
log_rzero_prior$expected_values$resize(1)
log_rzero_prior$expected_values[1]$value <- log_rzero_prior_mean
log_rzero_prior$log_sd$resize(1)
log_rzero_prior$log_sd[1]$value <- log(log_rzero_prior_sd)
log_rzero_prior$set_distribution_links(
"prior",
recruitment$log_rzero$get_id()
)
# Growth
growth <- FIMS:::initialize_growth(
parameters = parameters,
data = data_4_model
)
# Maturity
maturity <- FIMS:::initialize_maturity(
parameters = parameters,
data = data_4_model
)
# Population
population_module_ids <- c(
recruitment = recruitment$get_id(),
growth = growth$get_id(),
maturity = maturity$get_id(),
fleets = purrr::map(fleet, \(x) x$get_id())
)
population <- FIMS:::initialize_population(
parameters = parameters,
data = data_4_model,
# TODO: need to remove linked_ids from the function and add module_id to the
# parameters tibble
linked_ids = population_module_ids
)
# # Put a prior on natural mortality
# M_prior <- methods::new(DlnormDistribution)
# M_prior$expected_values$resize(1)
# M_prior$expected_values[1]$value <- inputs[["ctl"]][["MG_parms"]][
# grep("NatM_", row.names(inputs[["ctl"]][["MG_parms"]])),
# "PRIOR"
# ]
# M_prior$log_sd$resize(1)
# M_prior$log_sd[1]$value <- inputs[["ctl"]][["MG_parms"]][
# grep("NatM_", row.names(inputs[["ctl"]][["MG_parms"]])),
# "PR_SD"
# ]
# M_prior$set_distribution_links(
# "prior",
# population$log_M$get_id()
# )
# Put a prior on init_naa
init_naa_prior <- methods::new(DnormDistribution)
init_naa_prior$expected_values$resize(1)
init_naa_prior$expected_values[1]$value <- init_naa_prior_mean
init_naa_prior$log_sd$resize(1)
init_naa_prior$log_sd[1]$value <- init_naa_prior_sd
init_naa_prior$set_distribution_links(
"prior",
population$log_init_naa$get_id()
)
fims_model <- methods::new(CatchAtAge)
fims_model$AddPopulation(population$get_id())
# Initialize the model
CreateTMBModel()